-Search query
-Search result
Showing all 42 items for (author: guerra & m)

EMDB-18182: 
Closed conformation of the g-tubulin ring complex nucleating microtubules
Method: single particle / : Llorca O, Serna M

EMDB-18181: 
Early closed conformation of the g-tubulin ring complex
Method: single particle / : Llorca O, Serna M, Fernandez-Leiro R

PDB-8q62: 
Early closed conformation of the g-tubulin ring complex
Method: single particle / : Llorca O, Serna M, Fernandez-Leiro R

EMDB-28850: 
SARS-CoV-2 Gamma 6P Mut7 S + COVA309-3 Fab
Method: single particle / : Torres JL, Ward AB

EMDB-28851: 
SARS-CoV-2 Gamma 6P Mut7 S + COVA309-10 Fab
Method: single particle / : Torres JL, Ward AB

EMDB-28852: 
SARS-CoV-2 Omicron 6P S + COVA309-35 Fab
Method: single particle / : Torres JL, Ward AB
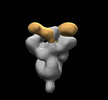
EMDB-28853: 
SARS-CoV-2 Gamma 6P Mut7 + S COVA309-38 Fab
Method: single particle / : Torres JL, Ward AB

EMDB-27177: 
sd1.040 Fab in complex with SARS-CoV-2 Spike 2P glycoprotein
Method: single particle / : Abernathy ME, Barnes CO

PDB-8d48: 
sd1.040 Fab in complex with SARS-CoV-2 Spike 2P glycoprotein
Method: single particle / : Abernathy ME, Barnes CO

EMDB-25412: 
Structure of the human COQ7:COQ9 complex by single-particle electron cryo-microscopy, unliganded state
Method: single particle / : Aydin H, Frost A

EMDB-25413: 
Structure of the NADH-bound human COQ7:COQ9 complex by single-particle electron cryo-microscopy
Method: single particle / : Aydin H, Frost A

PDB-7ssp: 
Structure of the human COQ7:COQ9 complex by single-particle electron cryo-microscopy, unliganded state
Method: single particle / : Aydin H, Frost A

PDB-7sss: 
Structure of the NADH-bound human COQ7:COQ9 complex by single-particle electron cryo-microscopy
Method: single particle / : Aydin H, Frost A
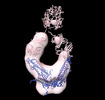
EMDB-26217: 
Negative stain EM map of COVA1-07 mAb bound to the S2 domain of SARS-CoV-2 S
Method: single particle / : Han J, Ward AB
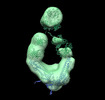
EMDB-26218: 
Negative stain EM map of COVA2-14 mAb bound to the S2 domain of SARS-CoV-2 S
Method: single particle / : Han J, Ward AB

EMDB-26219: 
Negative stain EM map of COVA2-18 mAb bound to the S2 domain of SARS-CoV-2 S
Method: single particle / : Han J, Ward AB
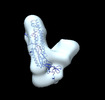
EMDB-26220: 
Negative stain EM map of the S2 domain of SARS-CoV-2 S
Method: single particle / : Han J, Ward AB

EMDB-13482: 
Half-vault structure
Method: single particle / : Guerra P, Gonzalez-Alamos M, Llauro A, Casanas A, Querol-Audi J, de Pablo P, Verdaguer N
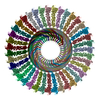
PDB-7pky: 
Half-vault structure
Method: single particle / : Guerra P, Gonzalez-Alamos M, Llauro A, Casanas A, Querol-Audi J, de Pablo P, Verdaguer N

EMDB-13478: 
Vault structure in primmed conformation
Method: single particle / : Guerra P, Gonzalez-Alamos M, Llauro A, Casanas A, Querol-Audi J, de Pablo P, Verdaguer N

EMDB-13483: 
Vault structure in committed conformation
Method: single particle / : Guerra P, Gonzalez-Alamos M, Llauro A, Casanas A, Querol-Audi J, de Pablo P, Verdaguer N

PDB-7pkr: 
Vault structure in primmed conformation
Method: single particle / : Guerra P, Gonzalez-Alamos M, Llauro A, Casanas A, Querol-Audi J, de Pablo P, Verdaguer N

PDB-7pkz: 
Vault structure in committed conformation
Method: single particle / : Guerra P, Gonzalez-Alamos M, Llauro A, Casanas A, Querol-Audi J, de Pablo P, Verdaguer N

PDB-7ko8: 
Cryo-EM structure of the mature and infective Mayaro virus
Method: single particle / : Riberio-Filho HV, Coimbra LD, Cassago A, Rocha RPF, Padilha ACM, Schatz M, van Heel MG, Portugal RV, Trivella DBB, de Oliveira PSL, Marques RE

EMDB-22961: 
Cryo-EM structure of the mature and infective Mayaro virus
Method: single particle / : Ribeiro Filho H, Coimbra LD, Cassago A, Rocha RFP, Padilha AC, Trivella DBB, Schatz M, Oliveira PSL, Portugal RV, Marques RE, van Heel M

EMDB-23502: 
Cryo-EM structure of the Pre3-1 20S proteasome core particle
Method: single particle / : Schnell HM, Walsh Jr RM

EMDB-23503: 
Cryo-EM structure of Pre-15S proteasome core particle assembly intermediate purified from Pre3-1 proteasome mutant (G34D)
Method: single particle / : Schnell HM, Walsh Jr RM

EMDB-23508: 
Cryo-EM structure of 13S proteasome core particle assembly intermediate purified from Pre3-1 proteasome mutant (G34D)
Method: single particle / : Schnell HM, Walsh Jr RM

PDB-7ls5: 
Cryo-EM structure of the Pre3-1 20S proteasome core particle
Method: single particle / : Schnell HM, Walsh Jr RM, Rawson S, Hanna JW

PDB-7ls6: 
Cryo-EM structure of Pre-15S proteasome core particle assembly intermediate purified from Pre3-1 proteasome mutant (G34D)
Method: single particle / : Schnell HM, Walsh Jr RM, Rawson S, Hanna JW

PDB-7lsx: 
Cryo-EM structure of 13S proteasome core particle assembly intermediate purified from Pre3-1 proteasome mutant (G34D)
Method: single particle / : Schnell HM, Walsh Jr RM, Rawson S, Hanna JW

EMDB-11236: 
Reconstruction of Tula virus surface glycoprotein lattice
Method: subtomogram averaging / : Stass R, Li S, Huiskonen JT

PDB-6zjm: 
Atomic model of Andes virus glycoprotein spike tetramer generated by fitting into a Tula virus reconstruction
Method: subtomogram averaging / : Stass R, Huiskonen JT, Rey F, Guardado-Calvo P
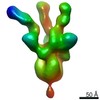
EMDB-22061: 
negative stain EM map of SARS-CoV-2 spike in complex with COVA2-15 Fab
Method: single particle / : Ward AB, Bangaru S, Torres JL

EMDB-22062: 
negative stain EM map of SARS-CoV-2 spike in complex with COVA1-22
Method: single particle / : Ward AB, Bangaru S, Torres JL

EMDB-22063: 
negative stain EM map of SARS-CoV-2 spike in complex with COVA2-07
Method: single particle / : Ward AB, Bangaru S, Torres JL

EMDB-22064: 
negative stain EM map of SARS-CoV-2 spike in complex with COVA2-39 Fab
Method: single particle / : Ward AB, Bangaru S, Torres JL

EMDB-22065: 
negative stain EM map of SARS-CoV-2 spike in complex with COVA2-04 Fab
Method: single particle / : Ward AB, Bangaru S, Torres JL

EMDB-22066: 
negative stain EM map of SARS-CoV-2 spike in complex with COVA1-12 Fab
Method: single particle / : Ward AB, Bangaru S, Torres JL

EMDB-0340: 
EM structure of the DNA wrapping in bacterial open transcription initiation complex
Method: single particle / : Florez-Ariza A, Cassago A, de Oliveira PSL, Guerra DG, Portugal RV

EMDB-4867: 
Sub-tomogram averaging of Tula virus glycoprotein spike
Method: subtomogram averaging / : Huiskonen JT, Stass R, Li S

EMDB-2101: 
Electron tomography of forming Hepatitis C virus-like particles in insect cells.
Method: electron tomography / : Badia-Martinez D, Peralta B, Andres G, Guerra M, Gil-Carton D, Abrescia NG
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
